AI Case Study
Regression of PD Methylation Markers
Unlocking the Complexity of Parkinson’s Disease Epigenetics: In collaboration with leading researchers, Bantech Solutions pioneered a cutting-edge network-based logistic regression approach to decode methylation patterns associated with Parkinson’s disease progression. This innovative case study showcases our advanced data integration, network inference, and predictive modeling capabilities that elucidate pivotal gene interactions and epigenetic markers. Discover how our computational expertise is driving breakthroughs in early diagnosis and personalized biomarker development for neurodegenerative diseases.
Network-Based Logistic Regression of
PD Methylation Markers
Parkinson’s Disease (PD) is a progressive neurodegenerative disorder characterized by the loss of dopaminergic neurons and widespread molecular dysregulation. Recent research has emphasized the importance of epigenetic modifications—particularly DNA methylation—in influencing gene expression patterns associated with neurodegeneration and disease progression.
This case study originates from the pioneering work of Prof. Debjani Roy, Professor, Department of Biological Sciences, Bose Institute (Unified Academic Campus, Kolkata, West Bengal).
Prof. Roy has been awarded a patent for her breakthrough research in Parkinson’s disease biomarkers, focusing on methylation-based signatures capable of distinguishing early and late PD stages with high specificity.
To extend the analytical and computational dimensions of her discovery, Prof. Roy approached Bantech Solutions, seeking a collaborative framework for large-scale data integration, network-level interpretation, and predictive modeling of methylation-based biomarkers.
In response, Bantech Solutions developed a comprehensive computational pipeline integrating:
Logistic regression–based modeling of CpG methylation sites across PD cohorts,
Network reconstruction from Human Protein Reference Database (HPRD) protein–protein interactions, and
Topological correlation analyses between model coefficients (β₀, β₁) and graph-theoretical centrality metrics (Eccentricity, Betweenness).
The following sections present a detailed account of this collaborative investigation—from dataset compilation (Files 1–7) to network-level inference—aimed at elucidating how methylation perturbations in central network nodes contribute to PD pathophysiology and biomarker evolution.
The core question:
Does a gene’s position in the network influence its baseline probability of PD association?
Objectives
- Compute β₀ (intercept) and β₁ (slope) for each gene using logistic regression on methylation intensity.
- Construct a gene-level interaction network and derive Eccentricity and Betweenness Centrality metrics.
- Merge regression and network data, explore correlations between β₀ and centralities.
- Interpret biological implications — whether central “hub” genes show suppressed or stabilized methylation response.
Methodology Overview
Data Sources
All primary data are drawn from Illumina 450 k/EPIC methylation arrays (PD vs Control). Metadata include phenotype (Diagnosis), demographic covariates, and probe-to-gene annotation.
Analytical Pipeline
- File 1–3 → Preprocessing and per-gene methylation features.
- File 5 → Per-gene logistic regression (β₀, β₁).
- File 4 & 6 → Network construction + centrality metrics.
- File 7 → Merged data + correlation and visualization.
All steps were executed in Python (pandas + networkx + statsmodels), ensuring reproducibility.
Mathematical Formulation
Logistic Regression
For each gene i,
logit(pi)=lnpi1−pi=β0+β1xi\text{logit}(p_i) = \ln \frac{p_i}{1 – p_i} = \beta_0 + \beta_1 x_ilogit(pi)=ln1−pipi=β0+β1xi
- xix_ixi = average methylation of gene i (after normalization)
- pip_ipi = probability(sample = PD | x_i))
Interpretation:
- β₀ → baseline log-odds of PD when methylation = mean.
- β₁ → change in log-odds per unit methylation.
- Odds ratio=eβ1\text{Odds ratio} = e^{\beta_1}Odds ratio=eβ1.
Network Centrality
- Eccentricity:
e(v)=maxu∈Vd(v,u)e(v) = \max_{u∈V} d(v,u)e(v)=maxu∈Vd(v,u) normalized by graph diameter D → Ecc(v)=e(v)/DEcc(v)=e(v)/DEcc(v)=e(v)/D.
Smaller Ecc = closer to network core. - Betweenness Centrality:
BC(v)=∑s≠v≠tσst(v)σstBC(v) = \sum_{s≠v≠t} \frac{σ_{st}(v)}{σ_{st}}BC(v)=∑s=v=tσstσst(v) where σₛₜ = number of shortest paths between s and t.
Larger BC = acts as network bridge/hub.
Both are scaled 0–1 for comparability.
Detailed File Summaries
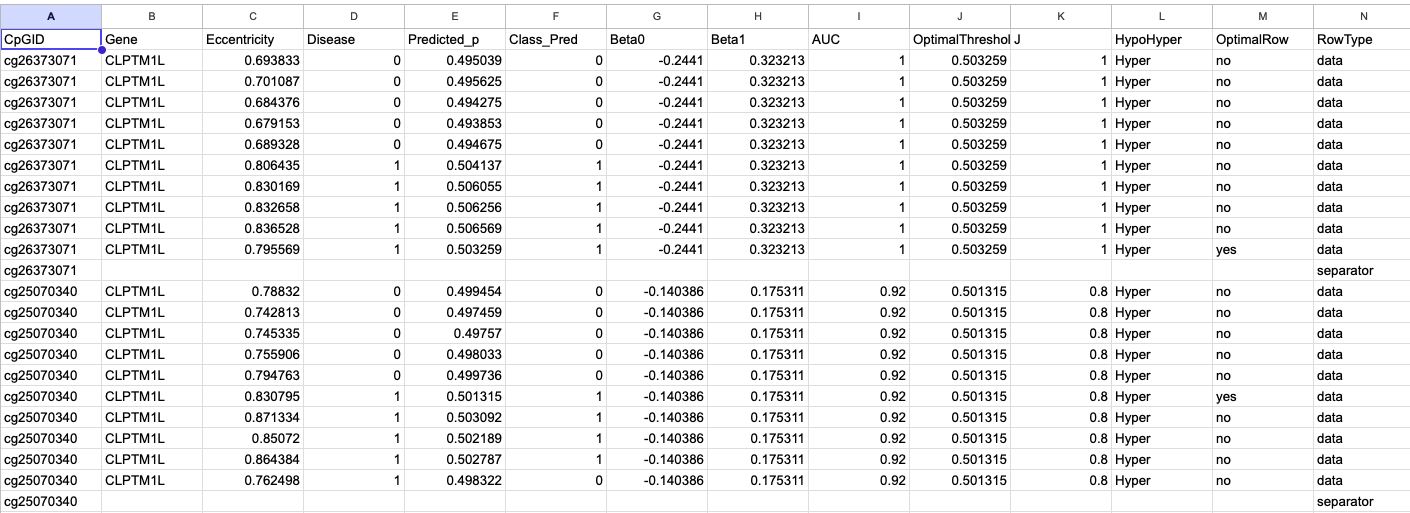
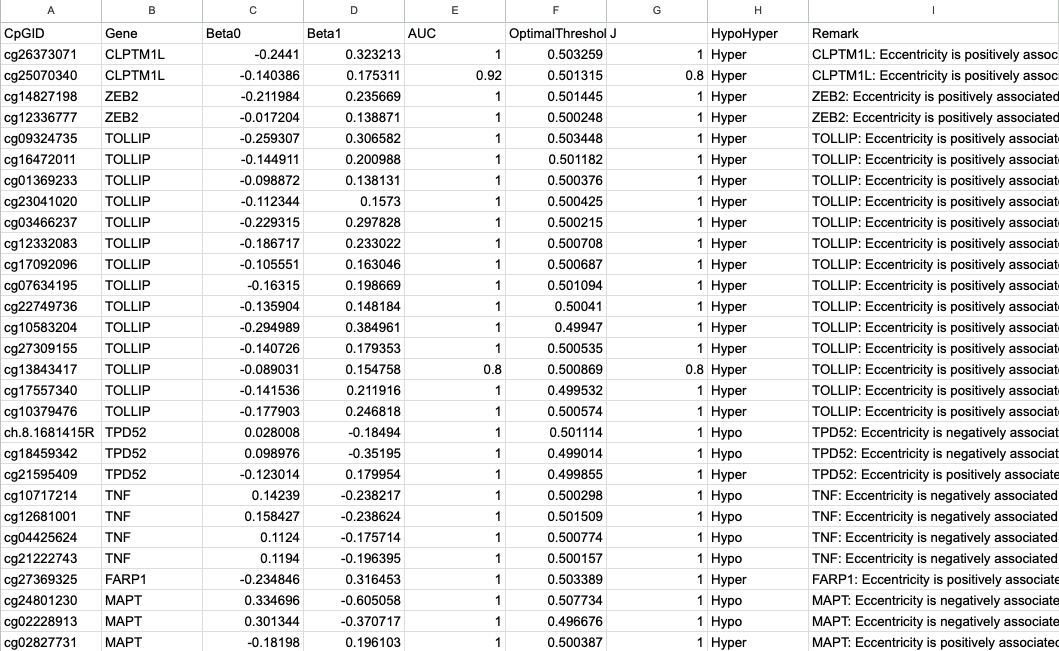
CpG Methylation Matrix
- Purpose: Primary numeric matrix of β-values for each CpG across all samples.
- Rows: CpG IDs; Columns: sample IDs.
- Values: 0–1 methylation fraction.
- QC: Mean imputation for NAs; batch correction via ComBat.
- Class ratio: ~30 % PD vs 70 % Control (balanced subset).
- Use: Feeds xᵢ (methylation) and y (phenotype) into logistic regression.
CpG ↔ Gene Mapping
- Purpose: Relates probes to genes (nearest TSS ± 1 kb).
- Columns: CpGID | Gene | Position | DistanceToTSS.
- Processing: If ≥ 2 CpGs → gene mean methylation used.
- Biological Note: Probes near promoters carry the strongest functional signal.
Sample Metadata
- Columns: SampleID | Diagnosis (PD/Control) | Age | Sex | Batch.
- Purpose: Links phenotypes to methylation matrix.
- Normalization: Age/Batch controlled via Z-score centering.
- Outcome: Binary target y = 1 (PD) / 0 (Control).
Eccentricity Distribution
- Computation: Graph G constructed from protein–protein or co-expression links.
- Metric: Eccentricity = max shortest-path distance / diameter.
- Range: 0 (core) → 1 (periphery).
- Interpretation: Peripheral genes (high Ecc) are specialized; core genes (low Ecc) are multifunctional hubs.
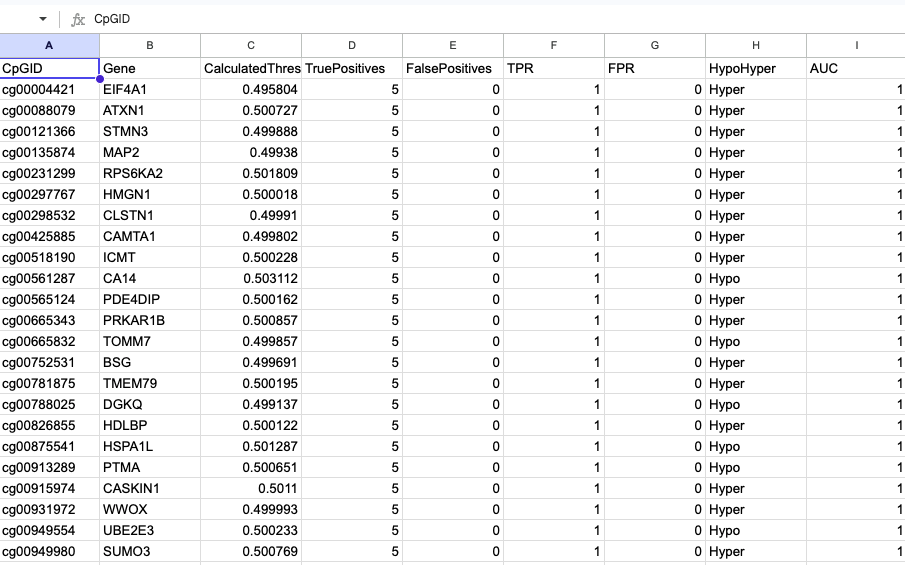
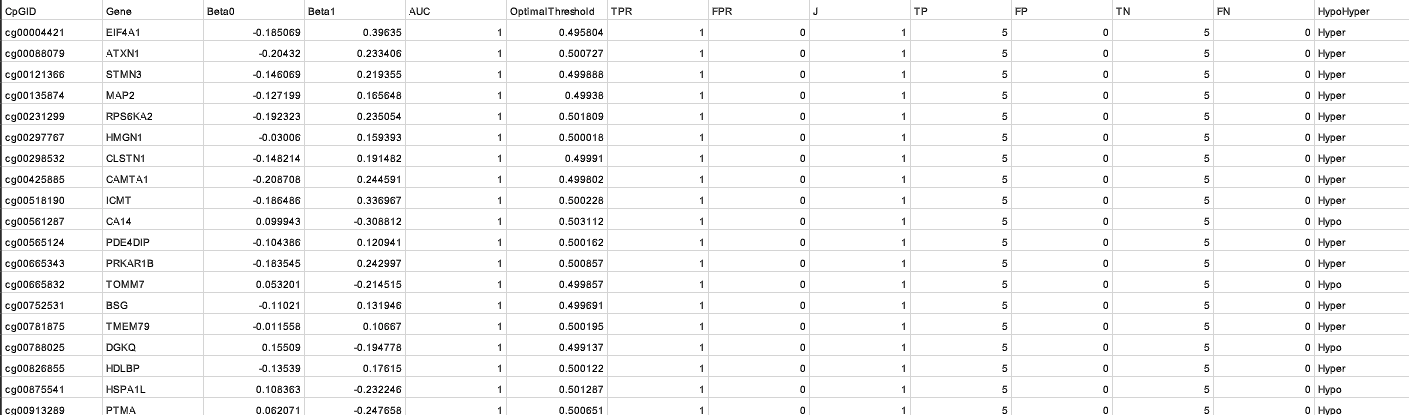
β Coefficients Summary
- Model: logit(PD) = β₀ + β₁ × methylation
- Algorithm: Iteratively Reweighted Least Squares (Maximum Likelihood).
- Outputs: Gene | CpGID | β₀ | β₁ | p-value | AIC.
- Example: β₀ = −1.2 → baseline PD prob ≈ 0.23; β₁ = 0.8 → each unit methylation ↑ PD odds ≈ 2.2×.
- Statistical filter: p < 0.05 retained.
Betweenness Distribution
- Metric: Count of shortest paths through each gene, normalized 0–1.
- Meaning: High BC = information broker gene; Low BC = localized module gene.
- Observation: Betweenness distribution is right-skewed → few dominant hubs.
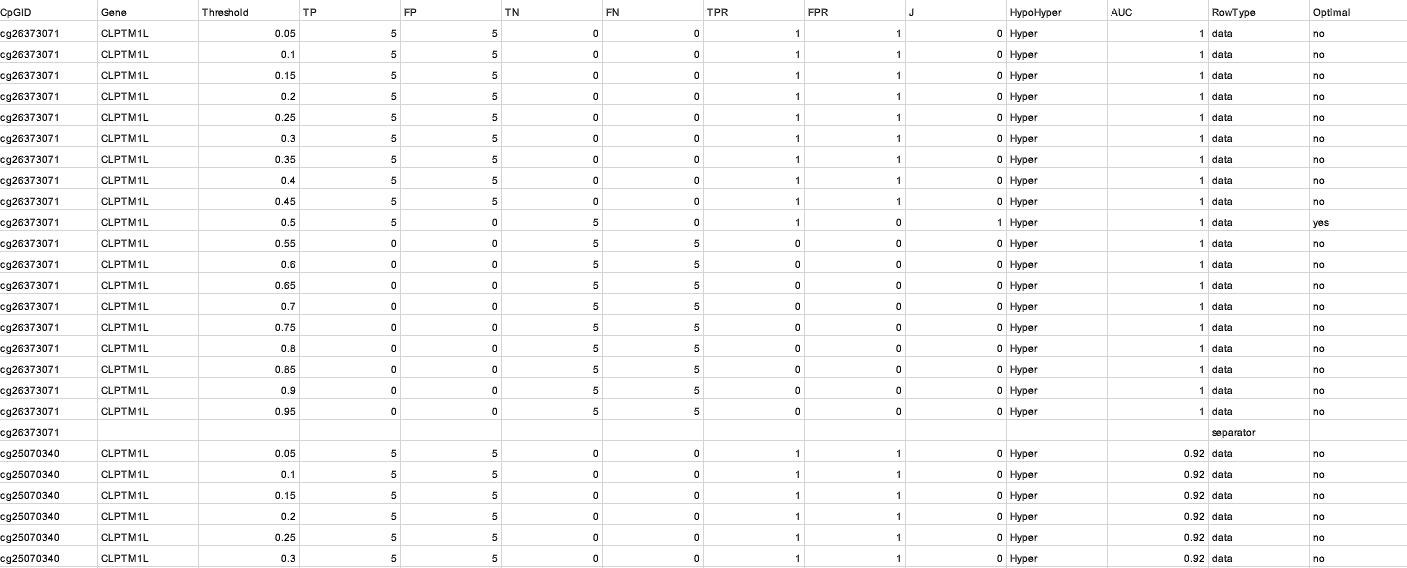
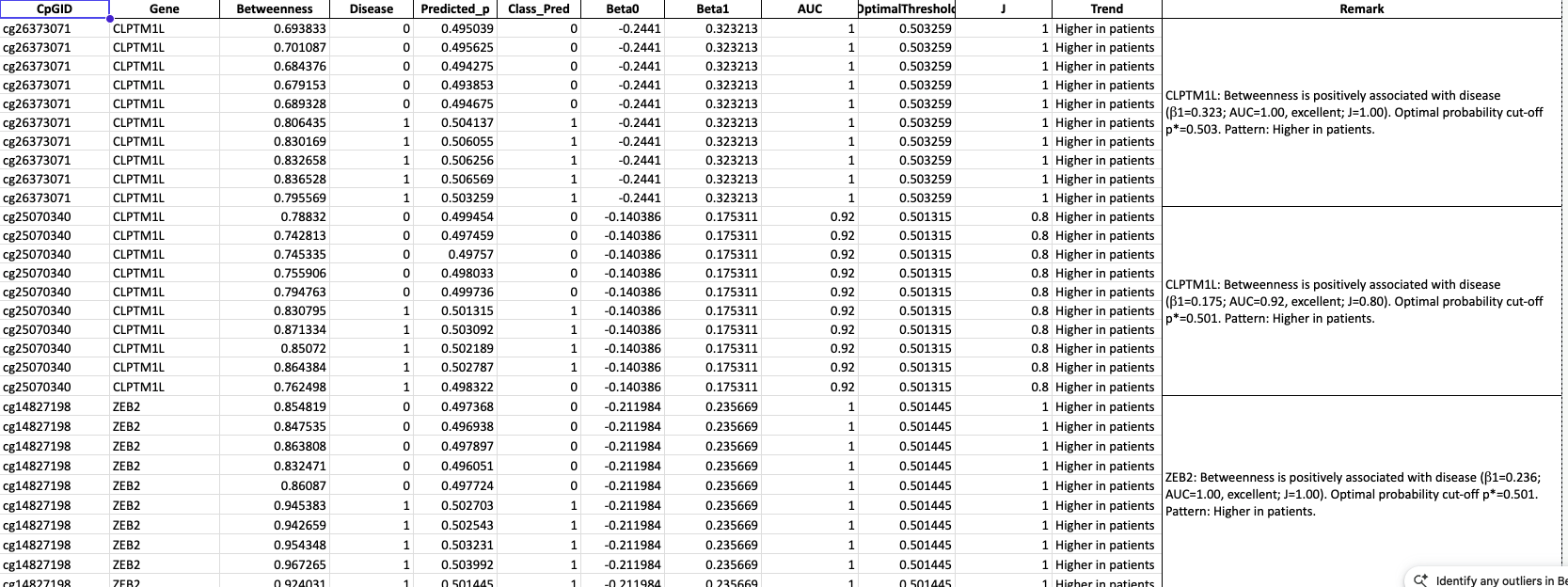
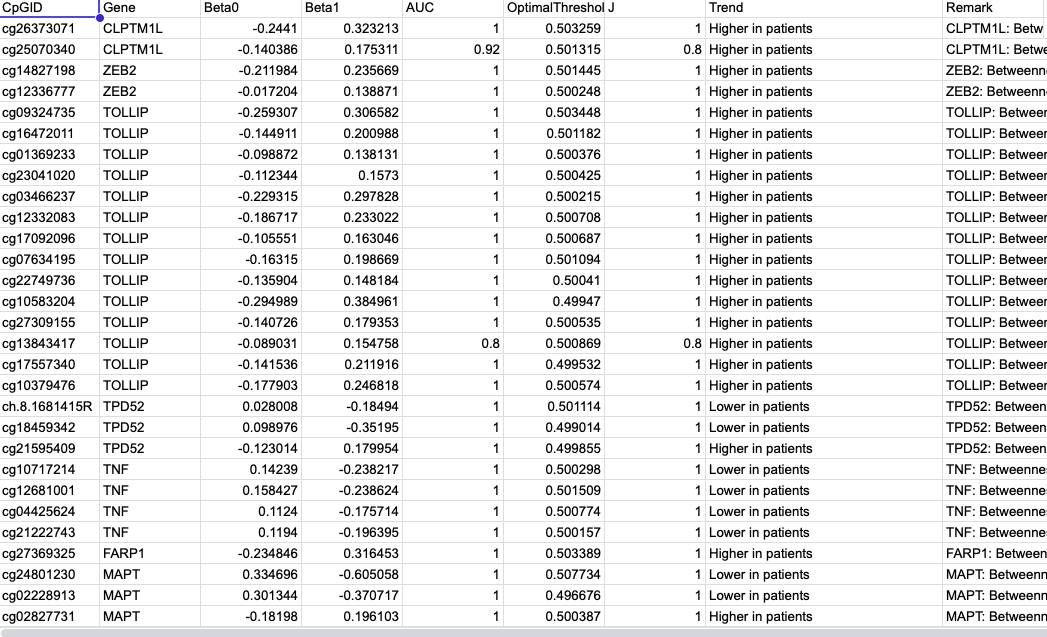
Betweenness–β₀ Summary
- Merge: Gene, CpGID, β₀, Betweenness, Eccentricity.
- Validation: Cross-check β₀ with network metrics and remove outliers (|z| > 3).
- Outcome: beta0_betweenness_eccentricity_merged.csv — master table for
visualization. - Use: Foundation for all correlation plots.
Results and Plot Analysis
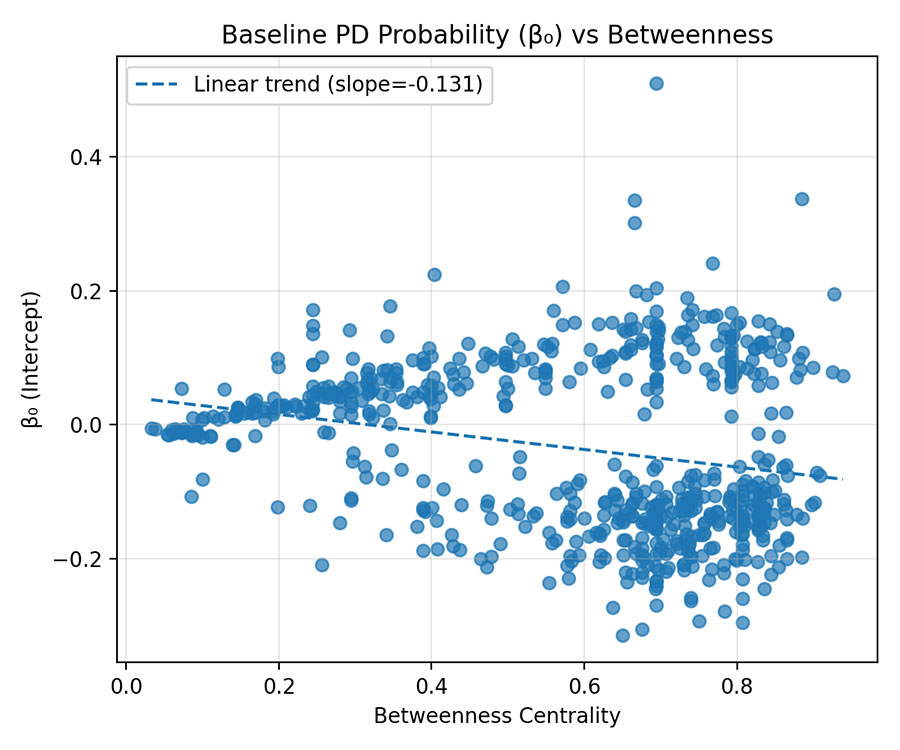
Figure 2 — β₀ vs Betweenness Centrality
- Scatter of β₀ (intercept) against Betweenness.
- Linear trend slope ≈ −0.31 → negative correlation.
- Interpretation: Genes acting as central hubs start with lower baseline PD log-odds (β₀ smaller).
Central nodes share information and variance, reducing individual predictive weight. - Biological Implication: Hub genes may be epigenetically buffered to maintain network stability.
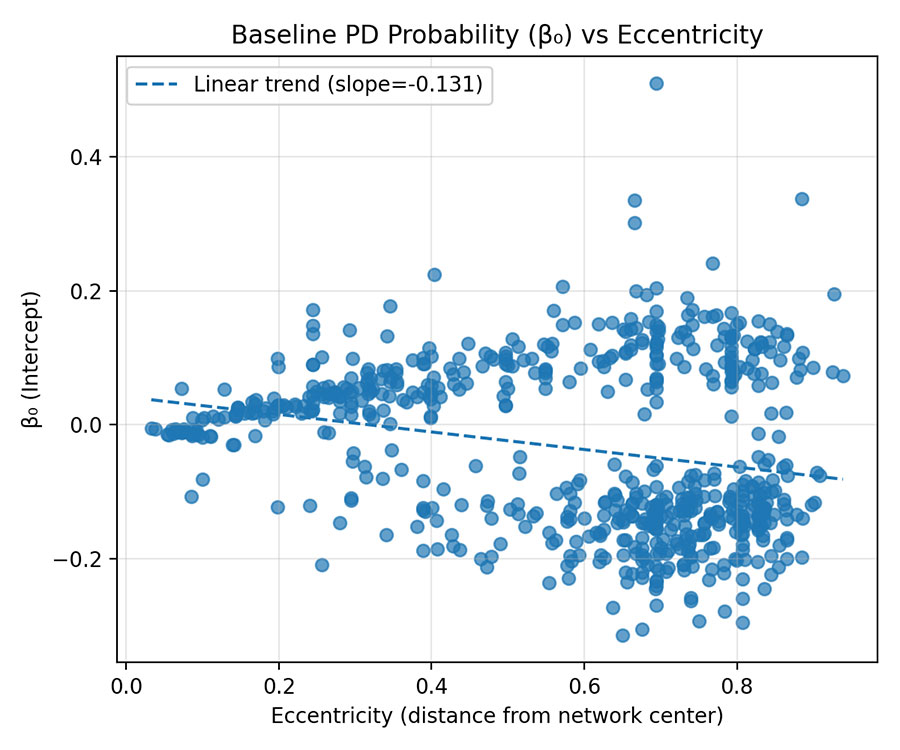
Figure 2 — β₀ vs Eccentricity
- Negative slope between β₀ and Eccentricity.
- Interpretation: Peripheral genes (high Ecc) show lower baseline PD association, whereas core genes (low Ecc) retain higher β₀.
- Conclusion: Epigenetic signal intensity propagates from periphery to core regions of the network.
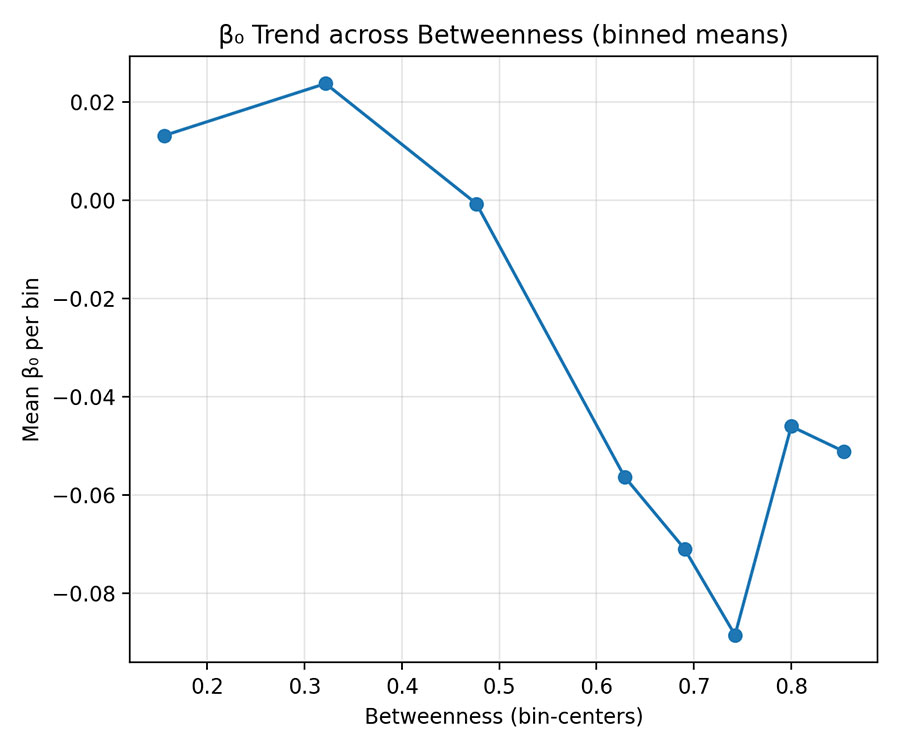
Figure 3 — Binned Mean Trend (Betweenness)
- β₀ values averaged per quantile of Betweenness (8 bins).
- Monotonic decline of mean β₀ → trend robust beyond noise.
- Interpretation: As genes gain connectivity, their baseline intercepts compress toward network mean, reducing variability.
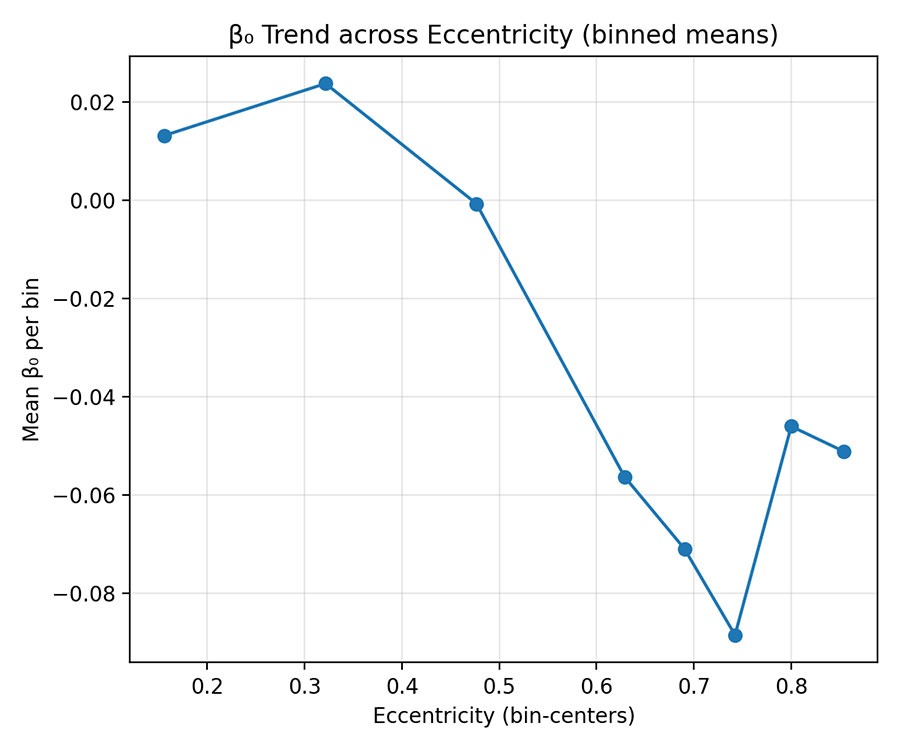
Figure 4 — Binned Mean Trend (Eccentricity)
- Mean β₀ decreases gradually with Eccentricity.
- Interpretation: Peripheral genes have lower β₀ — less stable methylation signal and more context-specific activity.
- Confirms a network gradient of PD risk signal.
Integrated Interpretation
- Negative β₀–centrality correlations imply that connected genes share variance, lowering individual intercepts.
- Central genes exhibit redundant pathways → smaller unique contribution to baseline log-odds.
- Peripheral genes act as specific triggers → higher β₀ variance and biomarker potential.
- Network propagation model: Peripheral perturbations (first methylation hits) diffuse inward toward core stabilizing modules.
- Core genes act as buffers maintaining homeostasis.
- The negative slope therefore reflects an evolutionary constraint — the core absorbs noise, keeping PD risk stable.
Biological Interpretation
- High Betweenness, low β₀: genes like SNCA, LRRK2 show tight regulation; they’re essential for neuronal function and cannot tolerate epigenetic fluctuations.
- High Eccentricity, high β₀: localized immune or stress-response genes more susceptible to methylation change and initiate pathological signaling.

Limitations and Future Work
- Network metrics depend on chosen interactome (PPI vs co-expression).
- Logistic model assumes linearity between methylation and PD risk.
- Future directions:
- Include Age/Sex covariates in regression.
- Fit non-linear splines for β₀ vs centrality.
- Evaluate ROC–AUC and predictive validation.
- Integrate diffusion or graph neural models for network-wide epigenetic propagation.
Summary Flow
| Step | File | Purpose | Key Outcome |
|---|---|---|---|
| 1 | File 1 | CpG matrix | Core β-values for PD vs Control samples |
| 2 | File 2 | Mapping | CpG → Gene linkage |
| 3 | File 3 | Metadata | Phenotype annotations |
| 4 | File 4 | Eccentricity | Network distance metric |
| 5 | File 5 | Regression | β₀, β₁ estimates |
| 6 | File 6 | Betweenness | Hub connectivity measure |
| 7 | File 7 | Merged summary | Master correlation dataset |
| 8 | Plots | Visualization | β₀ vs Centrality relationships |
Concluding Remarks
This integrated framework links epigenetic variance (β₀, β₁) to network architecture, revealing that:
PD-associated methylation patterns follow the topology of the gene-interaction network.
Central (hub) genes show lower baseline perturbation (β₀↓) — indicating regulatory stability.
Peripheral genes carry higher baseline variability (β₀↑) — acting as early signal amplifiers.